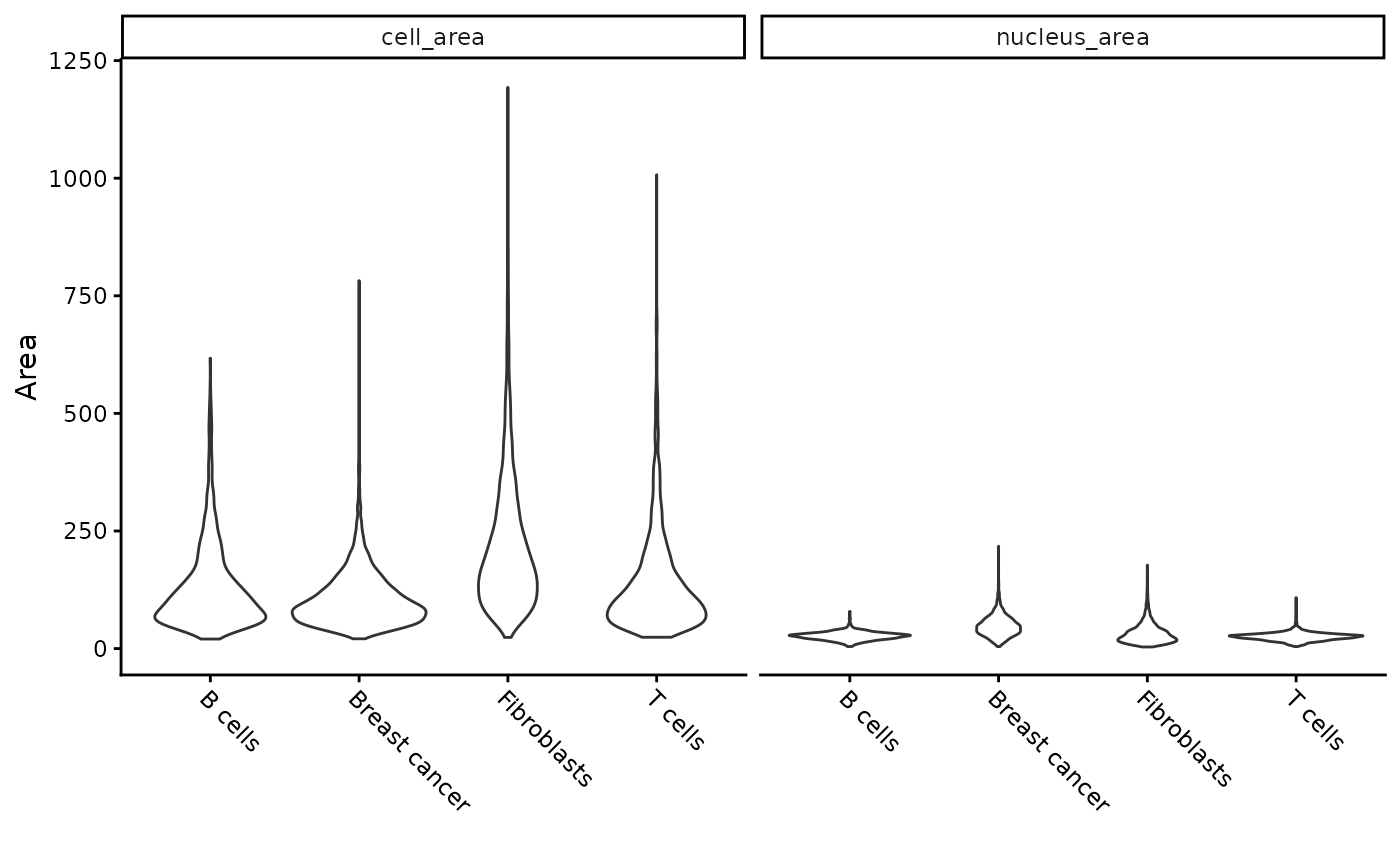
Violin plot using genes or cell data
Usage
plotViolin(
spe,
feature,
assay = "counts",
group.by = NULL,
type = c("raw", "log", "cpm", "logcpm"),
point = FALSE,
cols = NULL,
ncol = NULL,
pt.size = 0.3,
pt.alpha = 0.3,
pt.shape = ".",
ylab = "Expression",
xlab = NULL
)Arguments
- spe
A SpatialExperiment object.
- feature
can be a vector of gene names in rownames(spe), or column names in colData(spe) if those columns are numeric.
- assay
Name of assay to use for plotting feature.
- group.by
values to group plot by. Must be in colData of spe and must be either factor or character.
- type
Transformation to apply for the group/feature. Options are "raw" , "log", "cpm", "logcpm", or a function that accepts and returns a vector of the same length.
- point
Whether to plot points.
- cols
Colour palette for violins. Can be a vector of colours or a function that accepts an integer n and return n colours.
- ncol
Number of column if group.by is used.
- pt.size
Size of points.
- pt.alpha
Alpha of points between 0 and 1.
- pt.shape
Shape of points.
- ylab
Label for the y-axis.
- xlab
Label for the x-axis